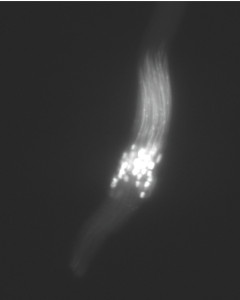
Segregation Distorter
Sex Ratio
Segregation Distorter (SD) is a well-studied selfish gene complex in D. melanogaster. In SD/SD+ heterozygous males, the SD chromosome is transmitted to 95% of the progeny by killing spermatids that have sensitive alleles of its target locus, Responder (Rsp)- a large satellite repeat near the centromere of chromosome 2. Despite knowing the molecular identity of both the distorter and the target, the mechanism behind distortion is unknown.
We use genomic, genetic, molecular and cytological approaches to study the mechanism behind segregation distortion and the effects of selfish genes on genome evolution and the genetic control of spermatogenesis. One hypothesis that we are testing is the possibility that small Rsp RNAs are affected by SD in the testis.
Modifiers of SD exist across the genome. Three enhancers of segregation distortion located on the 2nd chromosome ( E(SD), M(SD), and St(SD) ) are frequently held in linkage disequilibrium with the driver through inversions. Suppressors of segregation distortion on the 3rd ( Su(SD)3 ) and X ( Su(SD)X ) chromosomes segregate at high frequencies in natural populations. The molecular identities of these modifiers are currently unknown but discovering their identities promises to provide important insight into the mechanism behind segregation distortion. We are using genetic and association mapping methods to map the modifiers of SD.